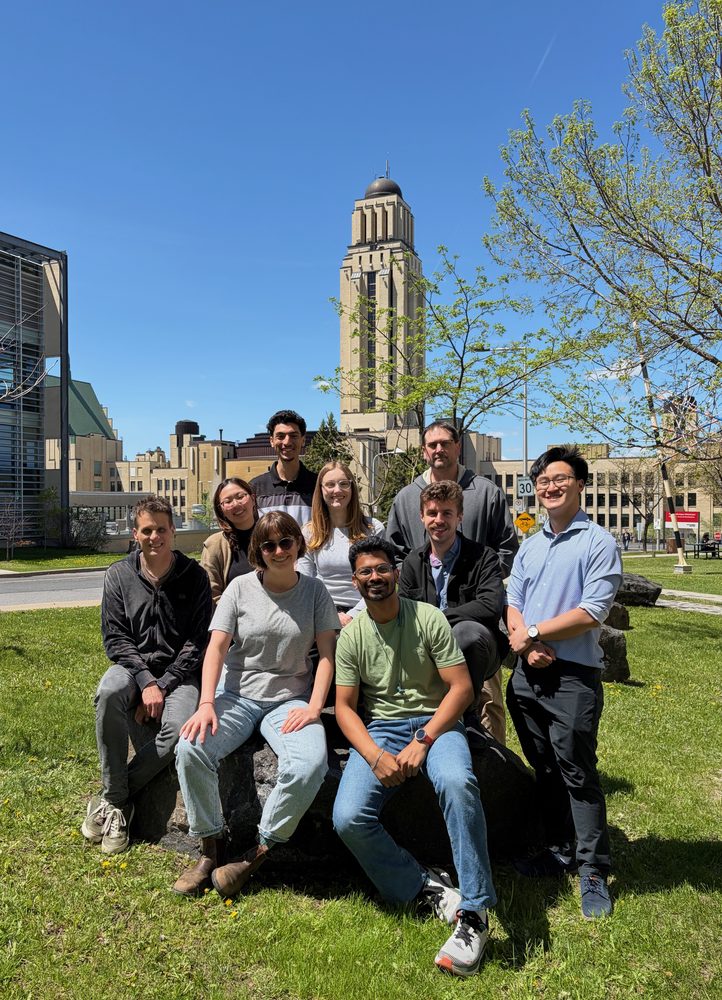
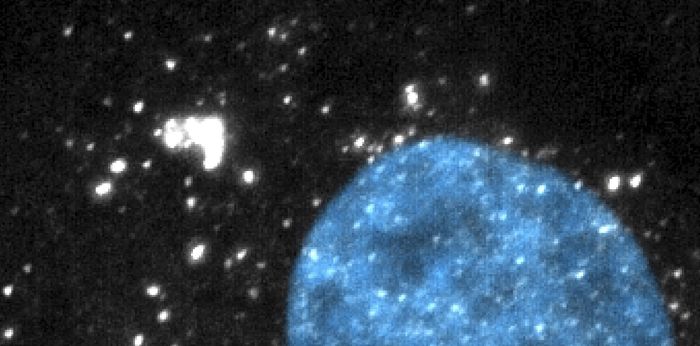
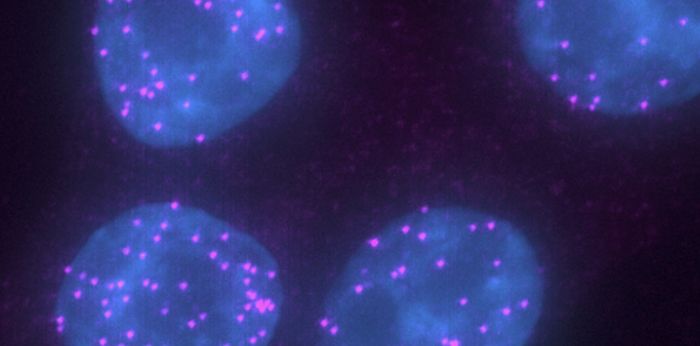
The Zenklusen lab studies how eukaryotic cells regulate the everyday choreography of gene expression: which RNAs leave the nucleus, which stay behind, and how their organization sets up the next steps of expression and decay. Most of the work happens at the level of single molecules and single cells, using yeast, mammalian and human cancer cell models.
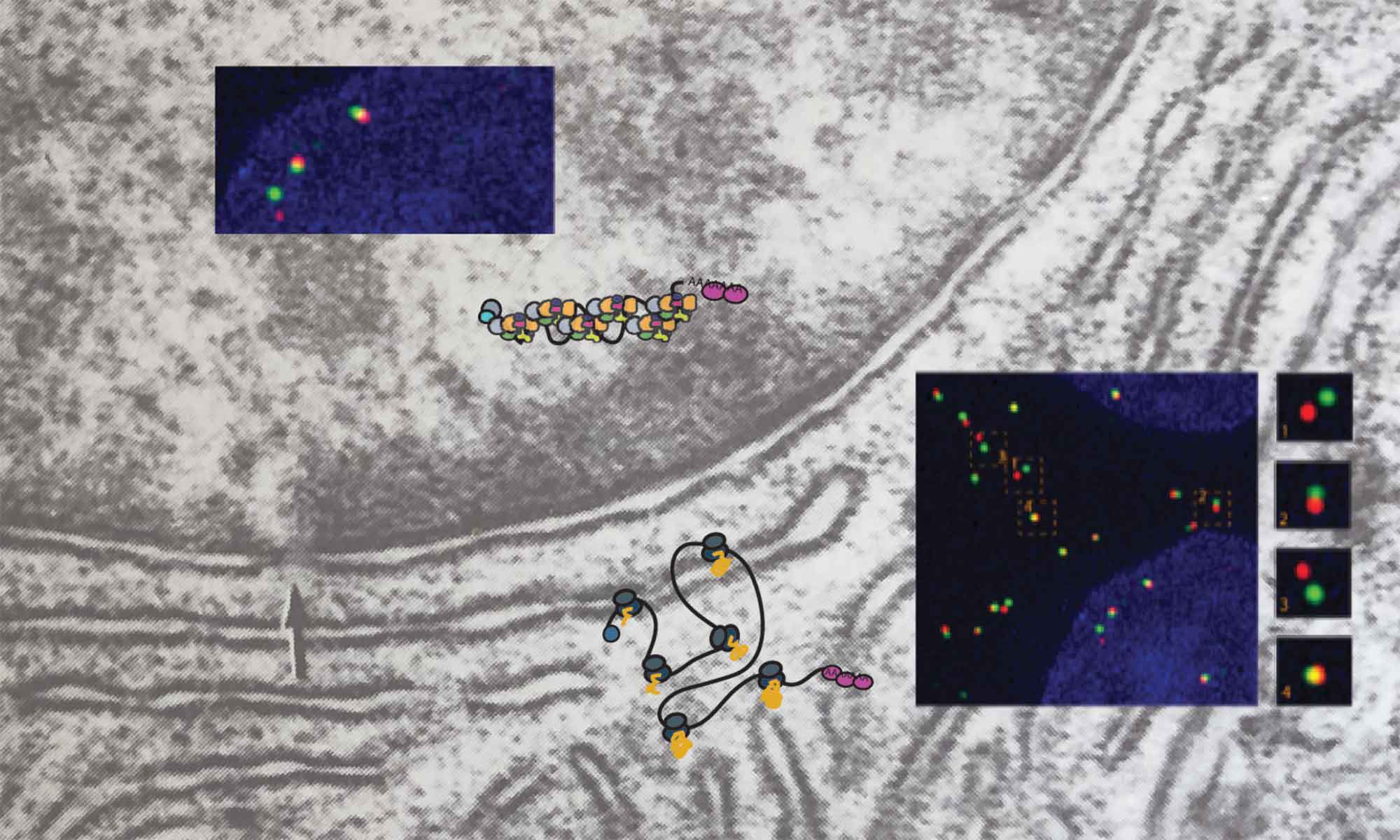
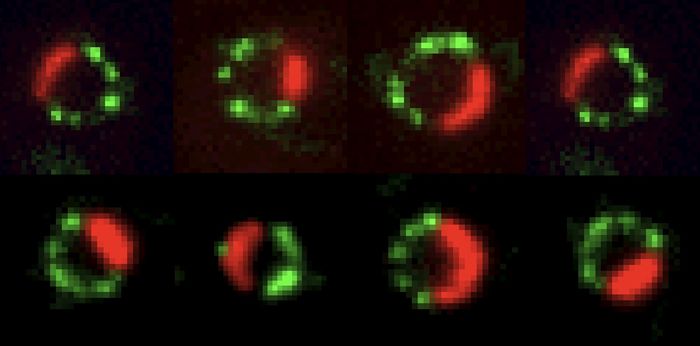
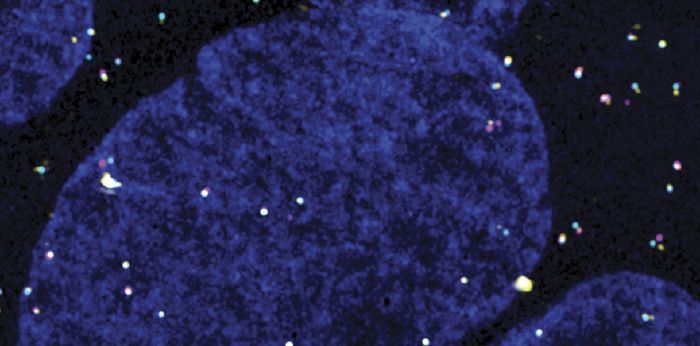
Over more than two decades, Daniel has helped develop the imaging tools that make this kind of biology possible — quantitative smFISH and MS2/PP7 live-cell imaging during his postdoc in Robert Singer’s lab, and 3D-SIM/STORM mRNP mapping more recently in Montreal — and applied them to a series of questions about RNA: how mRNAs are packaged into mRNPs — both those on their way to the cytoplasm and intronic RNAs before and after splicing, how those particles are selected by the nuclear basket of the nuclear pore, how non-coding RNAs tune transcription, and, most recently, how cells take care of repeat elements — whether from normally dormant LINE-1 or from Alu elements often hosted within introns.
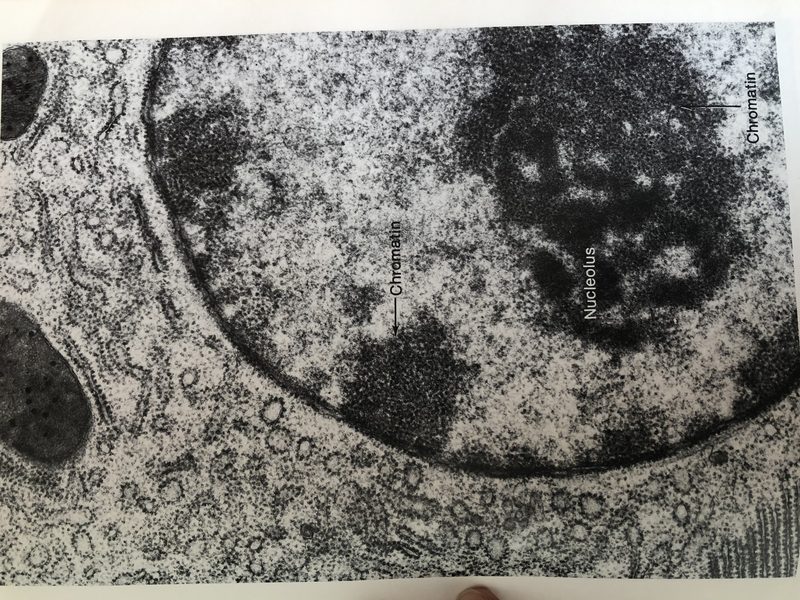
Current projects in the lab span four themes — RNA transport through the nuclear pore, (pre-)mRNP topology, ribosome biogenesis and nucleolar organization, LINE-1 biology, and post-splicing intron metabolism — held together by a shared commitment to quantitative, single-molecule readouts of cellular biology.